We have a new paper out in PLoS One that I think is particularly cool:
The paper is about species with big genomes – in particular species from the Dipteran family Tephritidae – a group of flies distantly (~150 million years) related to Drosophila. Tephritids are economically important – several species, most famously the Medfly Ceratitis capitata, are serious agricultural pests. But what’s interesting about them here is that they have big genomes – significantly bigger than your typical Drosophila species (450 to 850 Mb as compared to 175 Mb).
No one has yet sequenced a tephritid genome (although we are working on that now). But as part of a project to study the evolution of fly regulatory sequences (see also this paper), we sequenced a few developmentally important loci from each of these tephritid species. When the two (really fantastic) students who did this work (Brant Peterson and Emily Hare) got these sequences, they did the natural thing, and aligned them to each other. The result of this comparison is the heart of the paper:
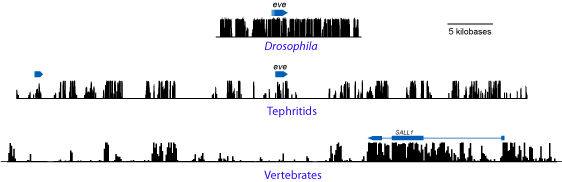
The three panels show plots of conservation (computed using PhastCons) across orthologous Drosophila and tephritid loci, and a vertebrate locus for comparison (the Drosophila and vertebrate analyses usesets of species that are at equivalent molecular distances as the tephritid species we analyzed – all plots are on the same scale – protein-coding genes are marked with blue boxes).
The point should be clear. The tephritid comparison looks much more like the vertebrate comparison than the Drosophila comparison.
Why is this important? Because for over a decade genome comparisons like those shown above have been successfully used to identify regulatory sequences in vertebrate genomes. Comparisons of the human, mouse, dog, chicken and fish genomes, for example, have identified thousands of discrete blocks of conserved non-coding sequences flanked by large stretches of rapidly evolving DNA – known generally as conserved non-coding sequences (CNSs). When assayed in mice, many of these CNSs turn out to be transcriptional enhancers that control gene expression during development.
The success of such methods in vertebrates inspired the sequencing of the Drosophila pseudoobscura genome – in the hope that comparisons of D. melanogaster and D. psuedoobscura genomes would lead to the rapid identification of enhancers across these genomes. But, outside of a few isolated cases, it didn’t happen. And the reason should be clear from the picture shown above: unlike in vertebrates, where islands of conservation stand out against a backdrop of rapidly evolving non-coding DNA, in Drosophila virtually all enhancer-sized chunks of non-coding DNA is conserved. Thus in Drosophila, while one can conclude that most non-coding DNA is functional, it is essentially impossible to determine where one regulatory sequence ends and the next begins.
Why the difference between vertebrates and Drosophila? It has been generally assumed that the difference arises from fundamental differences in the way that non-coding DNA is organized in vertebrates and invertebrates. But the tephritids show that this is not the case. The landscape of non-coding sequence conservation in tephritids looks like that of vertebrates – blocks of conserved DNA flanked by areas of rapidly evolving sequence. Drosophila looks different not because its non-coding sequences function in some less complex way than those of vertebrates – but rather because most of the rapidly evolving non-coding DNA found in other species has been deleted. The differences between Drosophila and vertebrates are real – but they are a product of Drosophila’s small genome, not of some essential difference in the structure structure and function of Drosophila and vertebrate genomes.
In part because Drosophila is such a high-profile invertebrate, many people extrapolated from vertebrate-Drosophila differences to conclude that there are fundamental differences in the organization of vertebrate and invertebrate genomes. This view has been immensely strengthened by a pervasive bias against sequencing large invertebrate genomes. Since most sequenced invertebrate genomes are small, the shared properties of small genomes – such as densely packed functional elements in non-coding DNA – have been mistakenly assumed to be shared properties of all invertebrate genomes.
Our data form the tephritids shows that this is not the case, as these large invertebrate genomes look – at least in this regard – like vertebrates. So, until we sequence more large invertebrate genomes, we are going to have a very biased view of the evolution of genome structure.
There’s also a much more practical consequence of our observations about tephritid genomes. As we show in our paper, many of the tephritid CNSs we identify function as enhancer in D. melanogaster embryos – this even though the species diverged around 150 million years ago. And, what’s more, we showed that despite extensive sequence divergence, we can map tephritid CNSs to the D. melanogaster genome, and that the tephritid CNSs drive expression patterns that are identical or very similar to those driven by the D. melanogaster sequences to which the tephritid CNSs map.
This immediately suggests that we can use comparisons of multiple tephritid species – and our mapping methods – to systematically annotate regulatory sequences in Drosophila genomes – something that comparisons of Drosophila species has not so far permitted.
There’s much more detail in the paper. Read it! And more importantly – comment on it, trash it, ask questions using the PLoS One commenting system. PLoS One is all about building a viable system of post-publication peer review – and we need you to participate to make it work.
3 Comments